Genomic imprinting is essential for mammalian embryonic development, and is an epigenetic phenomenon present in higher animals and plants. Dysregulation of the imprinted genes causes diabetes, cancer and neurological disorders. Understanding the expression regulation mechanism of imprinted genes can help us understand human individual development and disease etiology and treatment.
In a previous study, Prof. Li Xiajun’s group from SLST discovered that ZFP57 is a key regulator in genomic imprinting, and it maintains DNA methylation imprint at the Imprinting Control Regions (ICRs) of the target imprinted regions. They also found that Zfp57 mutants displayed maternal-zygotic effect, the first one identified in mammals. On January 27, 2021 Li’s group and their collaborators published a paper entitled “ZFP57 dictates allelic expression switch of target imprinted genes” in PNAS, revealing how ZFP57 controls the allelic expression switch of target imprinted genes.
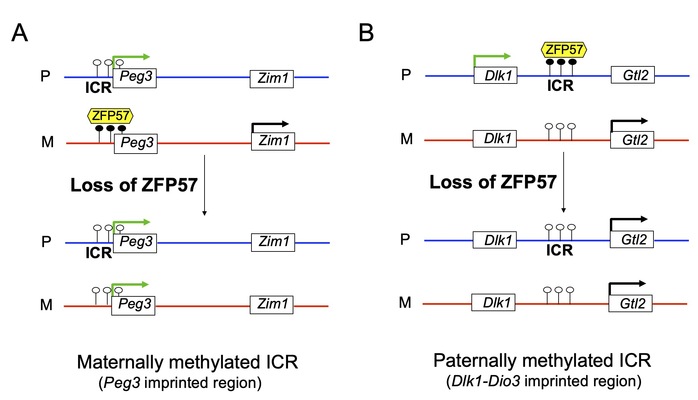
The regulated imprinted genes display parent-of-origin-dependent mono-allelic expression. Most imprinted genes are clustered and co-regulated by a cis-acting imprinting control region (ICR) in each imprinted region. In this study, Li’s group found that ZFP57 maintains DNA methylation imprint at most known ICRs. Mono-allelic expression of ZFP57 target imprinted genes is lost in Zfp57 mutant embryos. Instead, they become bi-allelic without ZFP57. The maternal alleles of some ZFP57 target imprinted genes will adopt similar expression patterns to those of their paternal counterparts upon loss of DNA methylation at the maternally methylated ICRs (e.g., Peg3 ICR) in Zfp57 mutant embryos (Fig. 1A). By contrast, the paternal alleles of some ZFP57 target imprinted genes will act like the maternal alleles when DNA methylation is lost at the paternally methylated ICRs (e.g., the IG-DMR of Dlk1-Dio3) in Zfp57 mutant embryos (Fig. 1B). This suggests there is an allelic expression switch of many ZFP57 target imprinted genes upon loss of DNA methylation at the ICRs without ZFP57.
In this study, ZFP57 can regulate the NOTCH signaling pathway in mouse embryos by impacting allelic expression as well as expression levels of a few regulators in the NOTCH signaling pathway based on RNA-seq analyses. These findings will help to uncover the underlying mechanisms in human diseases which are related to pathogenesis of imprinting. In addition, they may also provide insights into other mono-allelic expression phenomena that occur in the olfactory receptor and protocadherin genes in the nervous system, the antigen receptor genes in the immune system and X chromosome inactivation in females.
Collaborators are research groups from SLST, ShanghaiTech, Shanghai General Hospital, Center for Excellence in Molecular Cell Science, CAS, and Tsinghua University. Fourth year PhD candidate Jiang Weijun was the first author, Prof. Li Xiajun was one of the corresponding authors.
The link of the paper: https://doi.org/10.1073/pnas.2005377118


